
AlphaFold cracks the 50-year protein folding mystery, predicting 3D structures with atomic accuracy in minutes—revolutionizing biology, drug discovery, and structural bioinformatics.
Gather round, for I have a tale of the invisible architects of life—the Proteins. For fifty years, scientists have struggled to decipher the secret shapes these molecules fold into, knowing the sequence but blind to the structure. Only about 100,000 unique structures have been seen, a mere drop in the ocean compared to the billions of known sequences. But a new hero has emerged from the realm of deep learning, a neural network named AlphaFold, to bridge this vast gap. It entered the CASP14 arena, a blind test where the true structures were hidden from view.
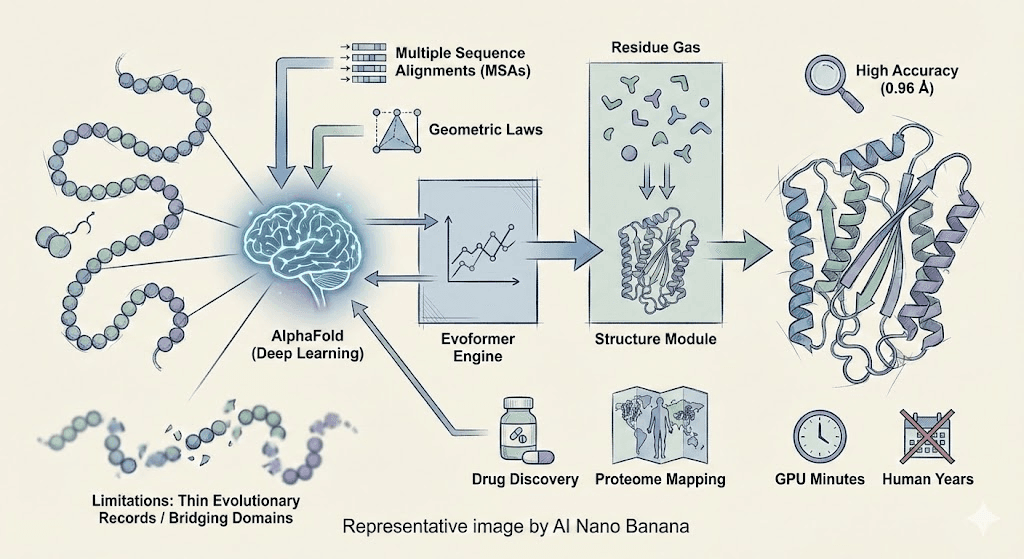
Unlike old methods that relied solely on slow physical simulations or sparse evolutionary hints, AlphaFold combined both paths. It listened to the whispers of evolutionary history through Multiple Sequence Alignments (MSAs) and the rigid laws of geometry. At its heart lay a powerful engine called the Evoformer. This engine allowed the network to exchange information constantly, treating the protein as a graph of relationships. Once the information was gathered, the Structure Module stepped in. It treated each amino acid as a floating rigid body—a 'residue gas'—free to rotate and translate. Through a process called 'recycling,' AlphaFold would refine its guess again and again, iterating until the shape was clear.
The true test came during the blind competition known as CASP14. AlphaFold faced off against the best existing methods, and the results were nothing short of magical. While the next best method stumbled with a median backbone accuracy of 2.8 Å, AlphaFold achieved a stunning 0.96 Å r.m.s.d.95. To understand this feat, picture a carbon atom, which is about 1.4 Å wide; AlphaFold was predicting positions with accuracy finer than the width of an atom itself. Even the side chains—the intricate details of the structure—were predicted with an all-atom accuracy of 1.5 Å, vastly outperforming the runner-up's 3.5 Å.
This breakthrough transforms the landscape of biology. Structures that once took months or years of painstaking effort to determine experimentally can now be predicted in GPU minutes or hours. This power opens the door to proteome-scale predictions, potentially mapping the entire human proteome. It has already proven its worth by aiding molecular replacement and interpreting cryogenic electron microscopy maps. By providing highly reliable per-residue confidence scores, AlphaFold gives scientists the courage to trust these computational models in real-world applications.
Yet, even the greatest heroes have their limits. AlphaFold relies heavily on the voices of the past. When the evolutionary record is thin—specifically when the median alignment depth drops below 30 sequences—its vision blurs. It also struggles with "bridging domains," proteins that only find their shape when embraced by a partner chain, relying heavily on heterotypic contacts.
Despite these limits, the gift AlphaFold offers is profound. It delivers structures in GPU minutes rather than human years, providing a reliable confidence score for every residue it predicts. It has already become a torchbearer for experimentalists, aiding in molecular replacement and interpreting maps from cryogenic electron microscopy. The seer has opened the door to the entire human proteome, turning the unknown into the known.
AlphaFold is a deep learning neural network developed by DeepMind that predicts the 3D structures of proteins from their amino acid sequences. It solved the decades-old protein folding problem by combining evolutionary information from multiple sequence alignments with geometric constraints, using a novel architecture called the Evoformer and a Structure Module that refines protein structures iteratively through a recycling process.
AlphaFold's accuracy is comparable to experimental techniques like X-ray crystallography and cryo-EM. In the CASP14 assessment, it achieved a median backbone accuracy of 0.96 Å r.m.s.d.95, which is finer than the width of a carbon atom (about 1.4 Å). Its all-atom predictions, including side chains, reached an accuracy of 1.5 Å, far surpassing previous methods.
AlphaFold struggles with proteins that have few evolutionary homologs (less than 30 sequences on average) and those requiring interaction with other chains to fold properly, such as 'bridging domains'. Despite these limits, it has revolutionized structural biology by enabling rapid, accurate structure prediction in minutes, aiding drug discovery, proteome-wide studies, molecular replacement, and interpretation of cryo-EM data with high per-residue confidence scores.
This article has been reviewed by a PhD-qualified expert to ensure scientific accuracy. While AI assists in making complex research accessible, all content is verified for factual correctness before publication.
AlphaFold: Revolutionizing Protein Structure Prediction
AlphaFold achieves atomic accuracy in protein structure prediction, transforming structural biology and bioinformatics.
How AlphaFold Solved the Protein Puzzle in Minutes?
Scientists built a smart computer named AlphaFold. It can guess the shapes of tiny body builders called proteins super fast!
How Scientists Count Tiny Protein LEGO Pieces in Your Body
Mass spectrometry helps scientists count proteins like LEGO blocks in your body. It’s like a super-smart scale for tiny building blocks!
No comments yet. Be the first to share your thoughts!
Get notified when we publish new articles. No spam, unsubscribe anytime.