
AlphaFold achieves atomic accuracy in protein structure prediction, transforming structural biology and bioinformatics.
Imagine being able to predict the 3D shape of any protein just from its genetic sequence—accurately, quickly, and without months of lab work. That’s exactly what AlphaFold, a groundbreaking AI system, has achieved. In a field where experimental methods like X-ray crystallography have dominated for decades, AlphaFold marks a paradigm shift by delivering computational predictions with atomic-level precision, often matching or rivaling real experimental structures.
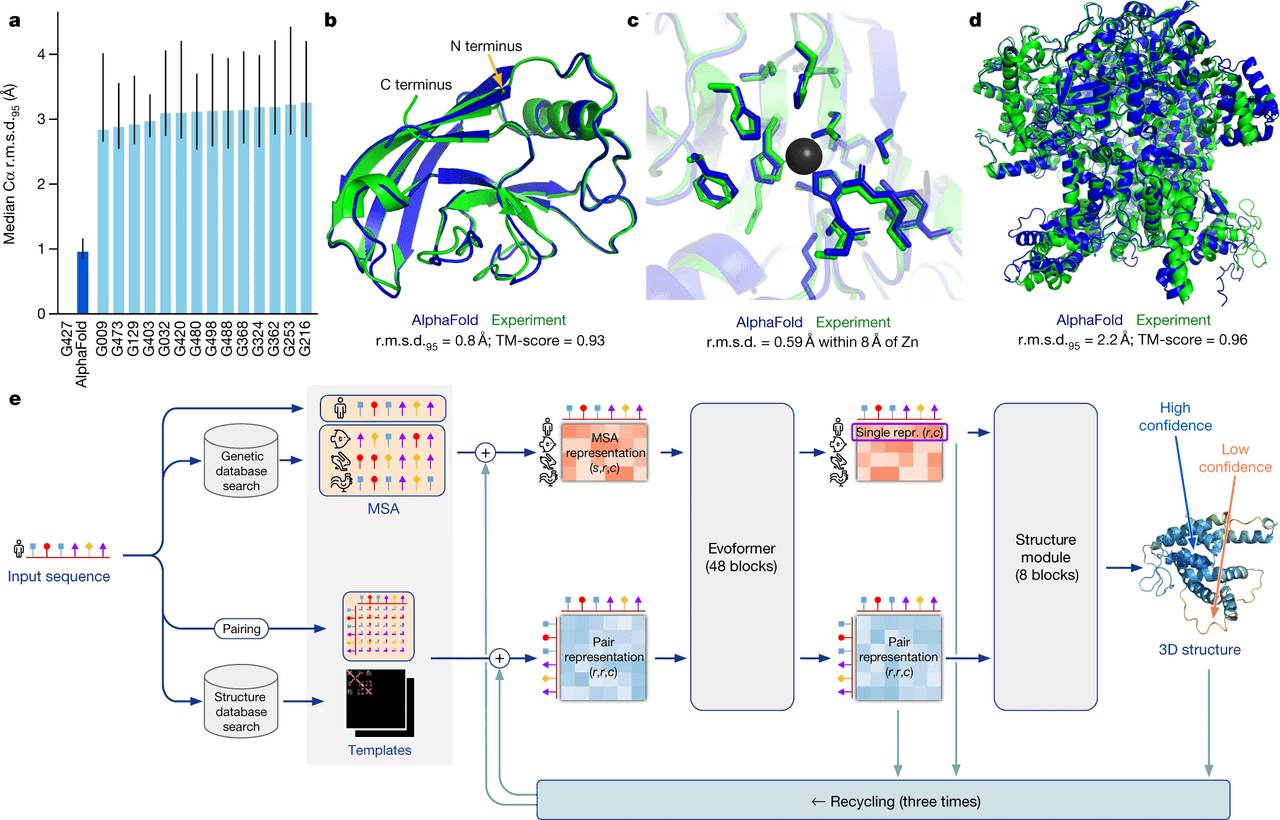
The team behind AlphaFold demonstrated that their model could predict protein structures with a median backbone accuracy of 0.96 Å r.m.s.d.95—a measure of how closely the predicted structure matches the real one. To put that in perspective, a carbon atom is about 1.4 Å wide, meaning AlphaFold’s predictions are nearly at the resolution of individual atoms. This performance was validated in the CASP14 competition, the gold standard for protein structure prediction, where AlphaFold outperformed all other methods by a huge margin—the next best had a median accuracy of 2.8 Å. Even more impressive, AlphaFold achieved this without relying on known structural templates, making it powerful for proteins with no evolutionary relatives in existing databases.
The model also produced highly accurate side-chain placements and correctly predicted complex features like zinc-binding sites—even though it wasn’t explicitly trained to detect metal ions. For a massive 2,180-residue protein with no structural homologues, AlphaFold accurately modeled both domain structures and how they pack together.
At its core, AlphaFold combines deep learning with biological and physical principles. The system uses two key inputs: the amino acid sequence of the protein and a multiple sequence alignment (MSA)—a collection of evolutionarily related sequences. These are processed through a novel neural network architecture called the Evoformer, which learns to extract evolutionary and spatial relationships between amino acids.
The Evoformer treats protein structure prediction as a graph inference problem, where residues are nodes and their interactions are edges. It uses attention mechanisms and a new “triangle multiplicative update” to ensure geometric consistency—like satisfying the triangle inequality in 3D space. This allows the network to build and refine a structural hypothesis layer by layer.
Then comes the structure module, which generates the actual 3D coordinates. It represents each residue as a local frame (rotation and translation), updating them iteratively using a technique called invariant point attention (IPA)—a geometrically aware attention mechanism that respects 3D symmetries. The model refines its prediction through multiple “recycling” steps, feeding its output back into the network for improvement, much like how humans revise drafts.
Crucially, AlphaFold doesn’t just predict structures—it also estimates its own confidence. The pLDDT score reliably predicts per-residue accuracy, and the pTM score estimates global fold correctness, enabling researchers to trust high-confidence predictions without experimental validation.
AlphaFold isn’t just a lab curiosity—it’s already transforming biology. It’s being used for molecular replacement in X-ray crystallography and to interpret cryo-EM maps, accelerating experimental workflows. Because it runs in minutes to hours on GPUs, it enables proteome-scale modeling—in a companion study, the team applied AlphaFold to the entire human proteome.
Yet, limitations remain. Accuracy drops when evolutionary data is sparse—specifically, when MSAs have fewer than ~30 sequences. The model also struggles with proteins whose shape depends heavily on interactions with other chains (heteromers), though it excels at predicting homomers, even highly intertwined ones.
Still, AlphaFold represents a quantum leap. By merging deep learning with physical and biological constraints, it has solved a 50-year-old grand challenge in biology. As genomic data explodes, AlphaFold bridges the gap between sequence and structure, paving the way for faster drug discovery, better enzyme design, and a deeper understanding of life at the molecular level.
AlphaFold is an artificial intelligence system developed to predict the 3D structure of proteins from their amino acid sequence with atomic-level accuracy. It represents a major breakthrough in structural biology because it solves a decades-old challenge—accurately determining protein structure—without requiring time-consuming experimental methods like X-ray crystallography or cryo-EM. Its predictions are so precise that they often match experimental results, enabling faster advances in drug discovery, disease understanding, and protein design.
AlphaFold combines deep learning with biological and physical principles using a sophisticated neural network architecture. It processes the protein's amino acid sequence and a multiple sequence alignment through the Evoformer, which models evolutionary and spatial relationships as a graph. The structure module then generates 3D coordinates using invariant point attention and iterative refinement, while confidence scores like pLDDT and pTM help assess prediction reliability, making the system both accurate and trustworthy.
AlphaFold performs best when there is sufficient evolutionary data—its accuracy decreases when few related sequences are available. It also has difficulty predicting structures of proteins that depend on interactions with other proteins (heteromers), though it excels at homomers. Despite these limits, it's being used to accelerate experimental methods like molecular replacement and cryo-EM analysis, and has enabled proteome-scale modeling, including the entire human proteome, transforming research in genomics, drug development, and synthetic biology.
This article has been reviewed by a PhD-qualified expert to ensure scientific accuracy. While AI assists in making complex research accessible, all content is verified for factual correctness before publication.
How AlphaFold Solved the Protein Puzzle in Minutes?
Scientists built a smart computer named AlphaFold. It can guess the shapes of tiny body builders called proteins super fast!
How Scientists Count Tiny Protein LEGO Pieces in Your Body
Mass spectrometry helps scientists count proteins like LEGO blocks in your body. It’s like a super-smart scale for tiny building blocks!
AlphaFold's Quest: Accurate Protein Structure Prediction
AlphaFold cracks the 50-year protein folding mystery, predicting 3D structures with atomic accuracy in minutes—revolutionizing biology, drug discovery, and structural bioinformatics.
No comments yet. Be the first to share your thoughts!
Get notified when we publish new articles. No spam, unsubscribe anytime.