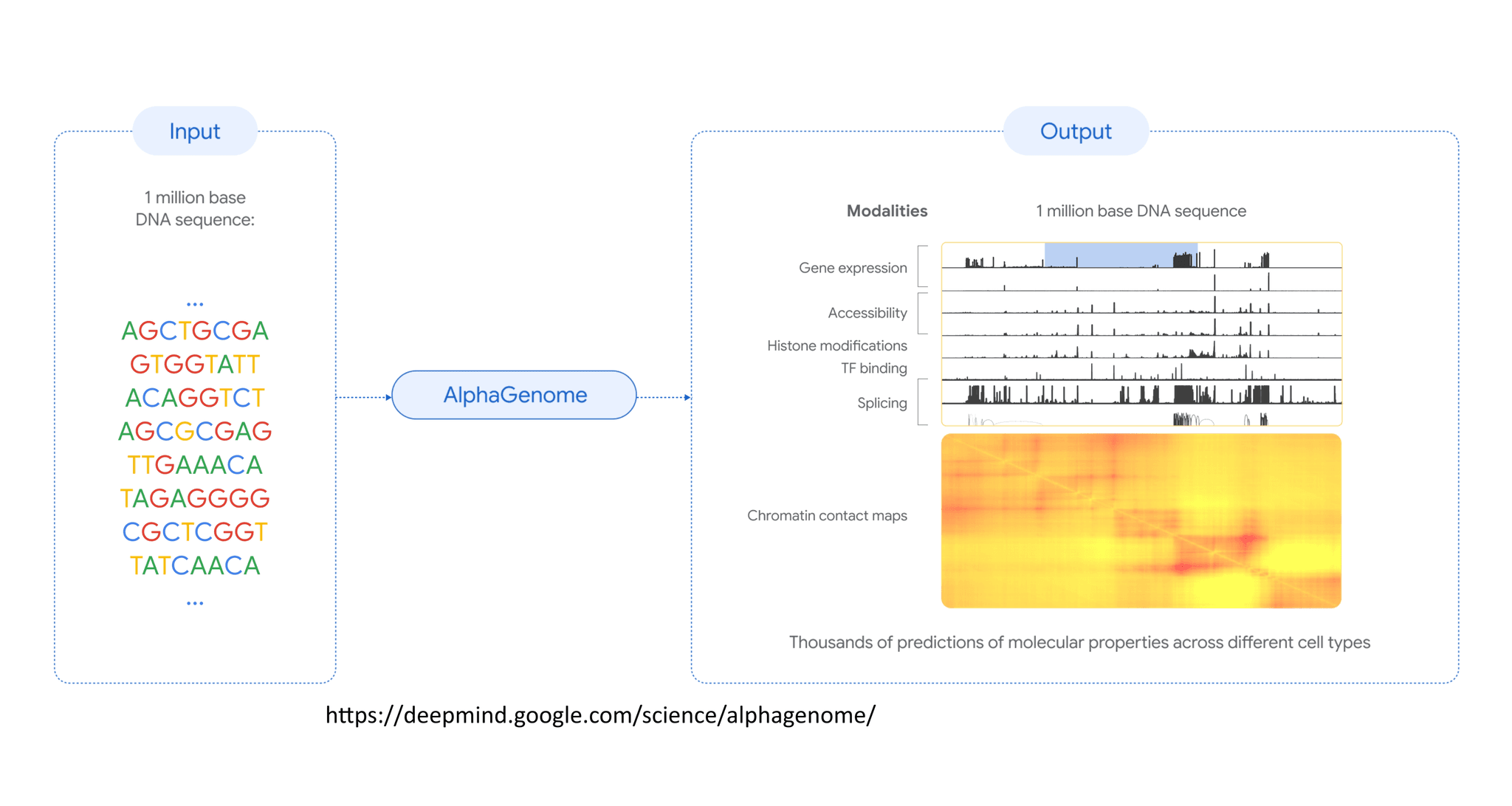


AlphaGenome is a new AI model from Google Deepmind team that analyzes massive 1-megabase chunks of DNA to predict how non-coding genetic regions function, achieving state-of-the-art performance in identifying disease-causing mutations and predicting gene regulation.
Imagine trying to read a book where 98% of the pages are written in a code we barely understand—that's non-coding DNA. But researchers from Google Deepmind just released a powerful new tool called AlphaGenome that is cracking this genetic regulatory code by processing massive chunks of DNA sequence to predict biological effects with incredible accuracy.
Here is the fascinating part: AlphaGenome doesn't just look at tiny snippets of DNA; it analyzes a massive 1 megabase (Mb) window at once. This allows it to predict thousands of functional genomic tracks, covering 11 diverse modalities like gene expression, chromatin accessibility, and splice site usage. When they tested it against existing models, AlphaGenome matched or exceeded the strongest available external models in 25 out of 26 evaluations of variant effect prediction. It even achieved state-of-the-art performance on 22 out of 24 genome track prediction tasks. For example, it showed a remarkable 14.7% relative improvement in predicting cell-type-specific gene expression changes compared to the previous top model, Borzoi. It even outperformed specialized models designed for single tasks, like beating ChromBPNet on accessibility predictions by 9.5% in profile accuracy. The model also demonstrated high fidelity in predicting splice junction counts, achieving a Pearson correlation of 0.74 in specific tasks.
Let me break down the mechanics for you. The model uses a U-Net-inspired architecture, which is excellent for processing complex data structures. It utilizes convolutional layers to spot local patterns—like specific transcription factor binding sites—and transformer blocks to understand long-range dependencies, such as how an enhancer far away might influence a gene promoter. Crucially, it processes this data at single-base-pair resolution, which is vital for seeing fine-scale features. To handle the massive computational load of processing 1 Mb of DNA, they used "sequence parallelism," splitting the work across multiple devices. They also employed a clever two-step training process involving "pretraining" and "distillation." First, they trained several "teacher" models on different parts of the genome. Then, a single "student" model learned to mimic the combined predictions of all those teachers using randomly augmented input sequences. This makes the final model highly efficient, taking less than a second to make a prediction on a modern GPU while maintaining high accuracy across all modalities.
The evidence suggests this could be a game-changer for understanding genetic diseases. Since the vast majority of genetic variation is non-coding, we need tools like this to figure out what those mutations actually do. The researchers demonstrated AlphaGenome's utility by accurately recapitulating the mechanisms of clinically relevant variants near the TAL1 oncogene. It successfully identified how specific mutations create a MYB motif that drives leukemia, illustrating its potential for clinical research. It’s particularly good at predicting splicing errors, which are a major cause of disease, offering a more comprehensive view than previous tools by simultaneously scoring splice sites, usage, and junctions. However, the researchers are honest about the limitations. The model still struggles with very distal elements (more than 100kb away) and capturing some tissue-specific nuances. It’s currently limited to human and mouse genomes and focuses mostly on protein-coding genes. Even so, AlphaGenome provides a much more unified and powerful foundation for analyzing the regulatory genome.
AlphaGenome is a powerful AI tool developed by Google DeepMind that helps decode non-coding DNA, which makes up about 98% of the human genome and has been poorly understood. By analyzing large segments of DNA—up to 1 megabase at a time—it can predict how genetic variants affect biological functions like gene expression and splicing, offering new insights into genetic regulation and disease.
AlphaGenome outperforms or matches existing models in nearly all evaluations. It achieved state-of-the-art results in 22 out of 24 genome track prediction tasks and showed a 14.7% improvement over the previous best model, Borzoi, in predicting cell-type-specific gene expression. It also surpassed specialized models like ChromBPNet by 9.5% in chromatin accessibility prediction and achieved high accuracy in splicing predictions with a Pearson correlation of 0.74.
AlphaGenome has significant potential for clinical research, such as identifying how non-coding mutations contribute to diseases like leukemia, as demonstrated with variants near the TAL1 oncogene. It excels at predicting splicing errors, a major cause of genetic disorders. However, it has limitations: it struggles with very distant regulatory elements (over 100kb away), may miss some tissue-specific effects, and currently only works on human and mouse genomes, focusing mainly on protein-coding genes.
This article has been reviewed by a PhD-qualified expert to ensure scientific accuracy. While AI assists in making complex research accessible, all content is verified for factual correctness before publication.
CRISPR-Cas9 Allele-Specific Trisomy 21 Chromosome Elimination in Human iPS Cells
A 2025 study used allele-specific CRISPR-Cas9 to eliminate the extra chromosome 21 from Down syndrome iPS cells, achieving a 30.6% karyotype correction rate in clones, outperforming non-specific methods and restoring gene expression linked to nervous system development.
CRISPR Gene-Edited Islet Cells Survive 12 Weeks Without Immunosuppression in Type 1 Diabetes Patient
Gene-edited islet cells survived 12 weeks in a type 1 diabetes patient with zero immunosuppression drugs, using CRISPR-Cas12b to achieve 100% HLA-II depletion.
Why Your Body Fights to Regain Lost Weight
Turns out, fat cells remember obesity in their DNA. Here's what that means for weight loss.
No comments yet. Be the first to share your thoughts!
Get notified when we publish new articles. No spam, unsubscribe anytime.