

VibeGen uses agentic AI to design de novo proteins based on vibrational motion. This 2026 study reveals that diverse protein folds can share similar dynamic behaviors, expanding protein engineering beyond static structures.
VibeGen is an agentic generative AI framework that designs de novo proteins based on specific vibrational motions rather than static structure. In this study, the system generated diverse, stable sequences that accurately matched prescribed low-frequency normal mode amplitudes. This matters because it proves that protein function can be engineered by targeting molecular motion directly.
Grab your coffee, because we are about to dive into a fascinating shift in how we engineer life's building blocks. The VibeGen generative AI protein dynamics design framework is challenging the status quo by treating motion as the primary design target, not just shape. Let me break this down.
The evidence suggests that focusing on dynamics rather than static geometry yields remarkably flexible results. In this study, the researchers demonstrated that multiple distinct protein folds can realize similar low-frequency dynamic behavior. This is a significant departure from traditional methods that target a specific shape. Instead of locking into one rigid structure, the VibeGen generative AI protein dynamics design system produced de novo sequences that exhibited no significant similarity to natural proteins. This expands the accessible protein space beyond evolutionary constraints. The study found that by incorporating a second agent to evaluate dynamic fidelity, the system raised the average correlation coefficient by filtering out designs that deviated from the target shape. This proves that designing for a single vibration shape does not necessarily fix the backbone or sequence, revealing a large degeneracy in sequence space.
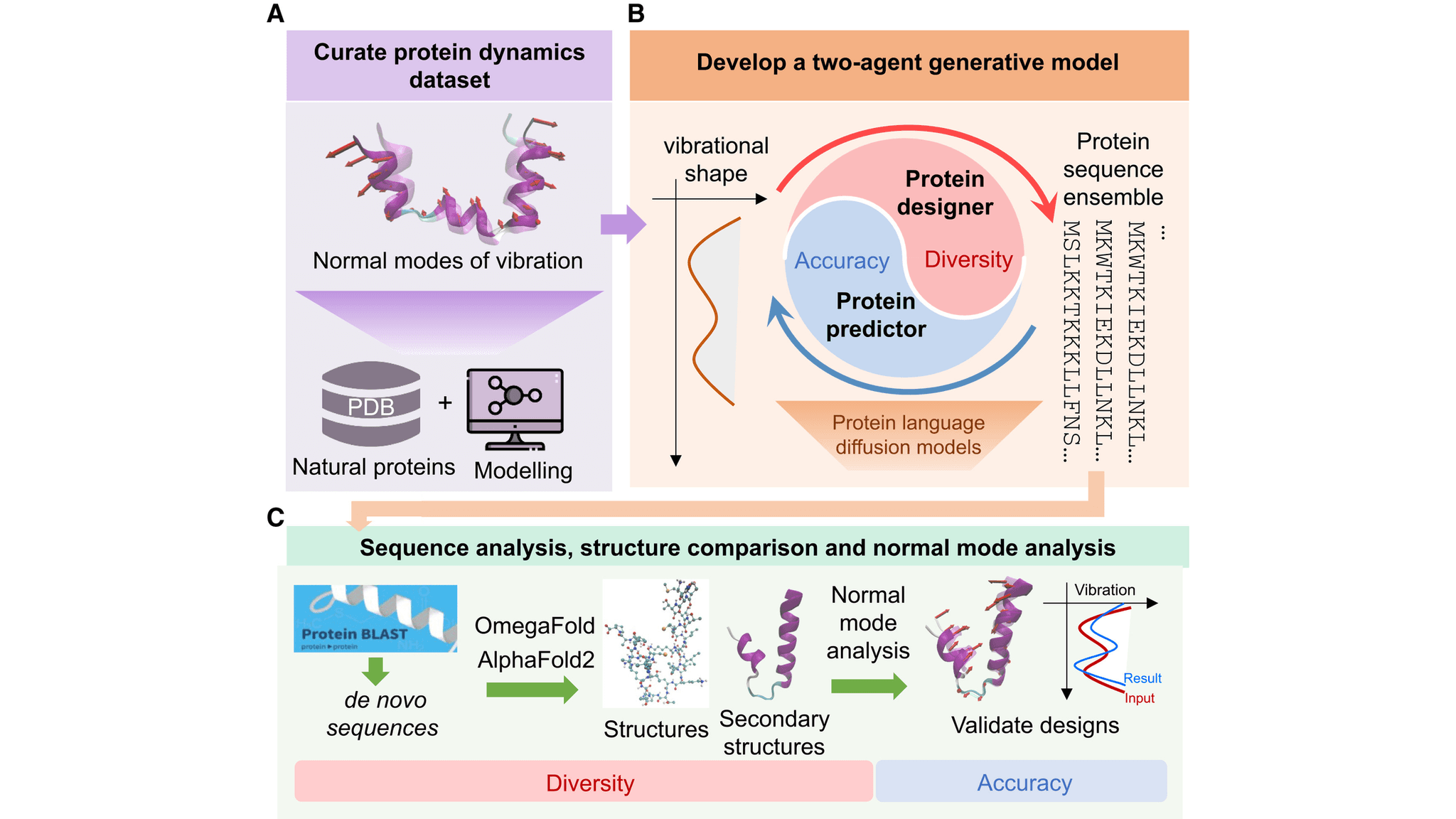
Here is the fascinating part. The system uses a dual-agent architecture comprising a Protein Designer and a Protein Predictor. The Protein Designer is trained to propose amino acid sequences based on given normal mode shapes, which represent the collective motions encoding how proteins bend and twist. The Protein Predictor then learns to predict the normal mode shapes for those sequences. At deployment, these two work collaboratively in an agentic loop to generate diverse yet accurate designs. This builds on earlier protein language diffusion models but adds a critical layer of physics-based validation. By coupling generation and evaluation, the model ensures the designed proteins accurately reproduce the prescribed normal mode amplitudes across the backbone. The researchers utilized full-atom molecular simulations to validate that the generated proteins adopt stable, functionally relevant structures while hitting the target dynamic markers.
The ability to directly program protein dynamics opens new opportunities to engineer enzymes with tailored catalytic motions and biomolecular materials with encoded mechanical responses. Similar to studies on AlphaFold, this work pushes the envelope toward a more complete understanding of protein functionality. However, the researchers acknowledge that challenges remain in capturing the full complexity of multi-scale dynamic phenomena. Comprehensive experimental validation using techniques like NMR spectroscopy or single-molecule FRET is crucial to confirm these in silico predictions. Additionally, while the framework successfully expanded sequence space, translating these insights into predictable biological outcomes in cellular contexts is the next hurdle. Future research may expand the model to include protein complexes or more detailed dynamics conditions like normal mode frequencies.
VibeGen uses a dual-agent system where a Protein Designer proposes sequences based on normal mode shapes and a Protein Predictor evaluates their dynamic accuracy. This agentic loop filters out designs that deviate from the target vibration, resulting in diverse, stable proteins that match prescribed low-frequency motions.
Protein functions like catalysis and signaling are governed by motion rather than just shape. By targeting low-frequency vibrational patterns, VibeGen can engineer proteins with specific functional movements, revealing that many distinct structural folds can share similar dynamic behaviors.
Current limitations include the need for experimental validation using techniques like NMR spectroscopy to confirm in silico predictions. The framework also faces challenges in capturing the full complexity of multi-scale dynamic phenomena and translating these insights into predictable biological outcomes in cellular contexts.
This article has been reviewed by a PhD-qualified expert to ensure scientific accuracy. While AI assists in making complex research accessible, all content is verified for factual correctness before publication.
The AI Hivemind: Why All Chatbots Sound the Same Now
You’ve noticed it too—AI responses are starting to blend together. Here’s why that’s dangerous.
Prompt Repetition Improves Non-Reasoning LLMs Without Added Latency
Repeating the input prompt improves accuracy across Gemini, GPT, Claude, and Deepseek models in 47 out of 70 benchmarks with zero losses and no added latency.
Anthropic's Assistant Axis: How LLM Persona Drift Causes Harmful AI Behavior
Researchers at Anthropic and Oxford identified a linear 'Assistant Axis' in LLM activation space that governs persona stability. Activation capping along this axis reduced harmful responses by nearly 60% without degrading model capabilities.
No comments yet. Be the first to share your thoughts!
Get notified when we publish new articles. No spam, unsubscribe anytime.